Gene Information
|
Gene Name
|
ETV1 |
|
Gene ID
|
2115
|
|
Gene Full Name
|
ETS variant transcription factor 1 |
|
Gene Alias
|
ER81 |
|
Transcripts
|
ENSG00000006468
|
|
Virus
|
HPV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Nucleoplasm; 
|
|
Membrane Info
|
Cancer-related genes, Disease related genes, Human disease related genes, Predicted intracellular proteins, Transcription factors |
|
Uniport_ID
|
P50549
|
|
HGNC ID
|
HGNC:3490
|
|
OMIM ID
|
600541 |
|
Summary
|
This gene encodes a member of the ETS (E twenty-six) family of transcription factors. The ETS proteins regulate many target genes that modulate biological processes like cell growth, angiogenesis, migration, proliferation and differentiation. All ETS proteins contain an ETS DNA-binding domain that binds to DNA sequences containing the consensus 5'-CGGA[AT]-3'. The protein encoded by this gene contains a conserved short acidic transactivation domain (TAD) in the N-terminal region, in addition to the ETS DNA-binding domain in the C-terminal region. This gene is involved in chromosomal translocations, which result in multiple fusion proteins including EWS-ETV1 in Ewing sarcoma and at least 10 ETV1 partners (see PMID: 19657377, Table 1) in prostate cancer. In addition to chromosomal rearrangement, this gene is overexpressed in prostate cancer, melanoma and gastrointestinal stromal tumor. Multiple alternatively spliced transcript variants encoding different isoforms have been identified. [provided by RefSeq, Jul 2016] |
Target gene [ETV1] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 5005469 |
chr7 |
74 |
84 |
5 |
163 |
View |
| 5005843 |
chr7 |
1 |
1 |
2 |
4 |
View |
| 5010743 |
chr7 |
0 |
2 |
3 |
5 |
View |
| 5011859 |
chr7 |
26 |
10 |
7 |
43 |
View |
Target gene [ETV1] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
C GSE183048
|
Chip-seq |
24 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE40774
|
Expression array |
134 |
Agilent-026652 Whole Human Genome Microarray 4x44K v2 (Probe Name version) |
|
GSE169622
|
Methylation profiling (Array) |
9 |
Infinium MethylationEPIC |
|
GSE181805
|
Expression array |
25 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE196215
|
RNA-seq |
8 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE140662
|
Expression array |
8 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE55542
|
Expression array |
36 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
C GSE143026
|
ATAC-seq;Chip-seq;RNA-seq |
30 |
Illumina HiSeq 2500 (Homo sapiens) |
|
E M TCGA_CESC
|
DNA methylation sequencing;RNA-seq |
288 |
TCGA |
|
GSE51993
|
Expression array |
48 |
Illumina Human v2 MicroRNA expression beadchip;Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE55550
|
Expression array |
155 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
GSE165883
|
RNA-seq |
20 |
Illumina NextSeq 500 (Homo sapiens) |
When the query gene is differentially changed in the dataset, a volcano/bar plot will be displayed.
> Dataset: TCGA_CESC - ETV1 expression across samples
|
Volcano Plot
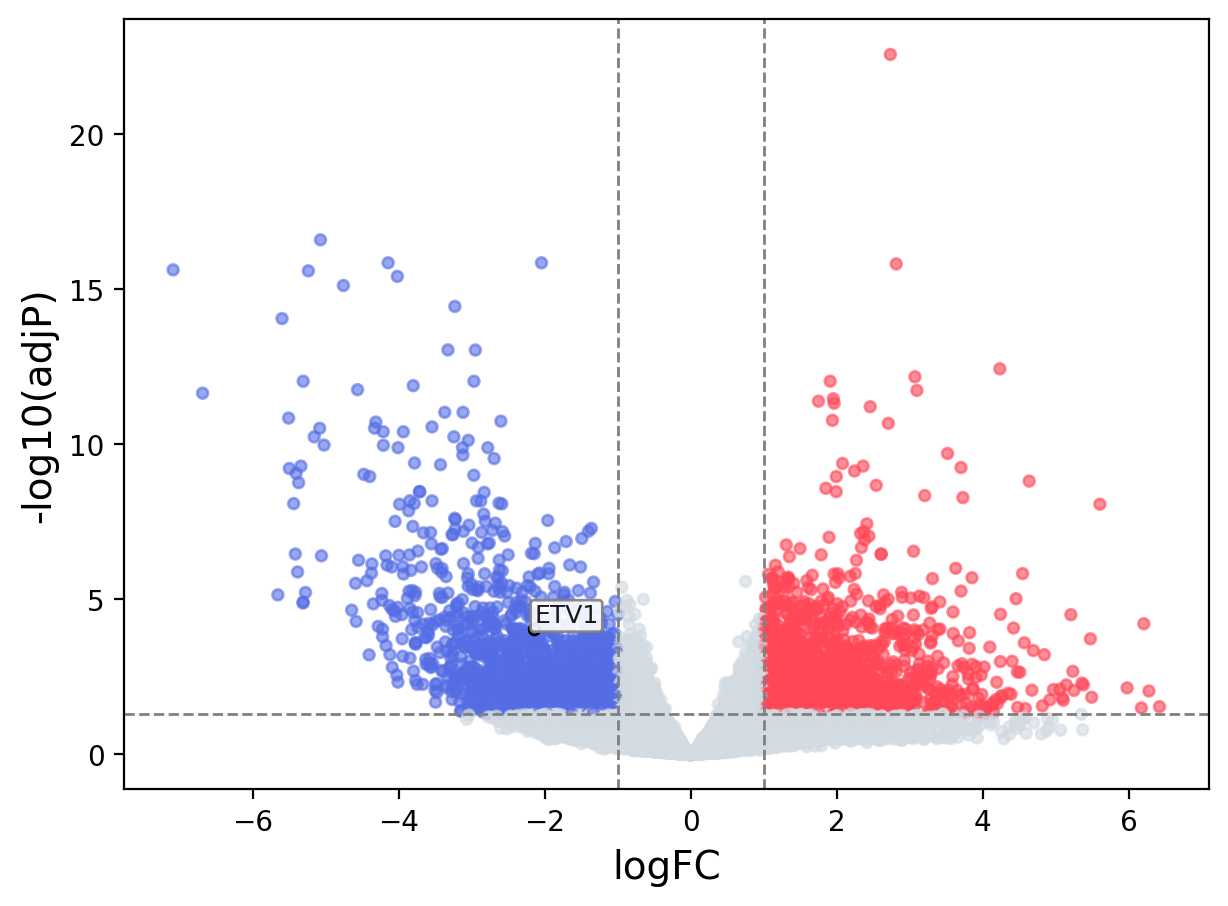
|
Bar Plot
|
|
|
 HPV Target gene Detail Information
HPV Target gene Detail Information