Expression profile of Mir7069 in omics data
| ID | GSE109165 |
| Title | Suppression of RGSz1 function optimizes the actions of opioid analgesics by mechanisms that involve the Wnt/β-catenin pathway |
| Organism | Mus musculus |
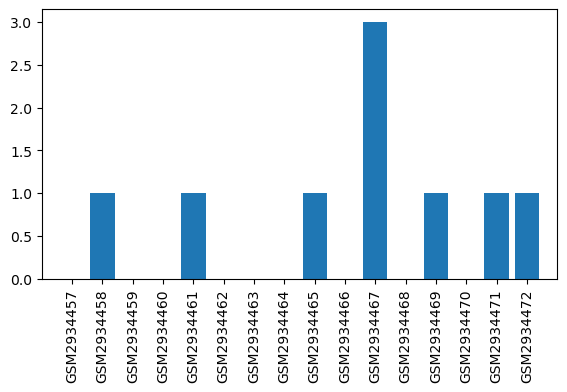
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2934457 | WT_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 0.0 |
| GSM2934458 | WT_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 1.0 |
| GSM2934459 | WT_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 0.0 |
| GSM2934460 | WT_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 0.0 |
| GSM2934461 | KO_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 1.0 |
| GSM2934462 | KO_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 0.0 |
| GSM2934463 | KO_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 0.0 |
| GSM2934464 | KO_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 0.0 |
| GSM2934465 | WT_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 1.0 |
| GSM2934466 | WT_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 0.0 |
| GSM2934467 | WT_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 3.0 |
| GSM2934468 | WT_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 0.0 |
| GSM2934469 | KO_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 1.0 |
| GSM2934470 | KO_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 0.0 |
| GSM2934471 | KO_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 1.0 |
| GSM2934472 | KO_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 1.0 |
| ID | GSE163828 |
| Title | Regulation of electro-acupuncture(EA) and astragalus membranaceus (AM) on genes in bone marrow |
| Organism | Mus musculus |
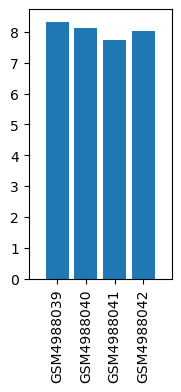
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4988039 | BH190126-1_control | tissue: bone marrow | 8.32 |
| GSM4988040 | BH190126-1_treatment of astragalus membranaceus | tissue: bone marrow | 8.12 |
| GSM4988041 | BH190126-1_cisplatin | tissue: bone marrow | 7.75 |
| GSM4988042 | BH190126-1_treatment of electro-acupuncture | tissue: bone marrow | 8.03 |
| ID | GSE223541 |
| Title | Oxycodone withdrawal induces HDAC1/2-dependent transcriptional maladaptations in the reward pathway in a murine model of peripheral nerve injury |
| Organism | Mus musculus |

| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6958536 | B1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sa | 0.0 |
| GSM6958537 | C1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958538 | F1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958539 | H1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |
| GSM6958540 | B2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 0.0 |
| GSM6958541 | C2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958542 | F2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958543 | H2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |
| GSM6958544 | B3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 0.0 |
| GSM6958545 | C3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958546 | F3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958547 | H3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |
| GSM6958548 | B4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 0.0 |
| GSM6958549 | C4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958550 | F4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958551 | H4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |

