GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0006325 | Chromatin organization |
| Process | GO:0006355 | Regulation of transcription, DNA-templated |
| Process | GO:0006357 | Regulation of transcription by RNA polymerase II |
| Process | GO:0006473 | Protein acetylation |
| Process | GO:0006475 | Internal protein amino acid acetylation |
| Process | GO:0006508 | Proteolysis |
| Process | GO:0006513 | Protein monoubiquitination |
| Process | GO:0006807 | Nitrogen compound metabolic process |
| Process | GO:0006996 | Organelle organization |
| Process | GO:0008152 | Metabolic process |
| Process | GO:0009889 | Regulation of biosynthetic process |
| Process | GO:0009891 | Positive regulation of biosynthetic process |
| Process | GO:0009893 | Positive regulation of metabolic process |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010390 | Histone monoubiquitination |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010556 | Regulation of macromolecule biosynthetic process |
| Process | GO:0010557 | Positive regulation of macromolecule biosynthetic process |
| Process | GO:0010604 | Positive regulation of macromolecule metabolic process |
| Process | GO:0016043 | Cellular component organization |
| Process | GO:0016567 | Protein ubiquitination |
| Process | GO:0016570 | Histone modification |
| Process | GO:0016573 | Histone acetylation |
| Process | GO:0016574 | Histone ubiquitination |
| Process | GO:0016578 | Histone deubiquitination |
| Process | GO:0016579 | Protein deubiquitination |
| Process | GO:0018193 | Peptidyl-amino acid modification |
| Process | GO:0018205 | Peptidyl-lysine modification |
| Process | GO:0018393 | Internal peptidyl-lysine acetylation |
| Process | GO:0018394 | Peptidyl-lysine acetylation |
| Process | GO:0019219 | Regulation of nucleobase-containing compound metabolic process |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0019538 | Protein metabolic process |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0031325 | Positive regulation of cellular metabolic process |
| Process | GO:0031326 | Regulation of cellular biosynthetic process |
| Process | GO:0031328 | Positive regulation of cellular biosynthetic process |
| Process | GO:0032446 | Protein modification by small protein conjugation |
| Process | GO:0035520 | Monoubiquitinated protein deubiquitination |
| Process | GO:0035521 | Monoubiquitinated histone deubiquitination |
| Process | GO:0035522 | Monoubiquitinated histone H2A deubiquitination |
| Process | GO:0036211 | Protein modification process |
| Process | GO:0043170 | Macromolecule metabolic process |
| Process | GO:0043412 | Macromolecule modification |
| Process | GO:0043543 | Protein acylation |
| Process | GO:0043966 | Histone H3 acetylation |
| Process | GO:0044238 | Primary metabolic process |
| Process | GO:0045893 | Positive regulation of transcription, DNA-templated |
| Process | GO:0045935 | Positive regulation of nucleobase-containing compound metabolic process |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048522 | Positive regulation of cellular process |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051173 | Positive regulation of nitrogen compound metabolic process |
| Process | GO:0051252 | Regulation of RNA metabolic process |
| Process | GO:0051254 | Positive regulation of RNA metabolic process |
| Process | GO:0051276 | Chromosome organization |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0070646 | Protein modification by small protein removal |
| Process | GO:0070647 | Protein modification by small protein conjugation or removal |
| Process | GO:0071704 | Organic substance metabolic process |
| Process | GO:0071840 | Cellular component organization or biogenesis |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:1901564 | Organonitrogen compound metabolic process |
| Process | GO:1902680 | Positive regulation of RNA biosynthetic process |
| Process | GO:1903506 | Regulation of nucleic acid-templated transcription |
| Process | GO:1903508 | Positive regulation of nucleic acid-templated transcription |
| Process | GO:2001141 | Regulation of RNA biosynthetic process |
| Component | GO:0000123 | Histone acetyltransferase complex |
| Component | GO:0000124 | SAGA complex |
| Component | GO:0005622 | Intracellular anatomical structure |
| Component | GO:0005634 | Nucleus |
| Component | GO:0005654 | Nucleoplasm |
| Component | GO:0031248 | Protein acetyltransferase complex |
| Component | GO:0031974 | Membrane-enclosed lumen |
| Component | GO:0031981 | Nuclear lumen |
| Component | GO:0032991 | Protein-containing complex |
| Component | GO:0043226 | Organelle |
| Component | GO:0043227 | Membrane-bounded organelle |
| Component | GO:0043229 | Intracellular organelle |
| Component | GO:0043231 | Intracellular membrane-bounded organelle |
| Component | GO:0043233 | Organelle lumen |
| Component | GO:0070013 | Intracellular organelle lumen |
| Component | GO:0070461 | SAGA-type complex |
| Component | GO:0071819 | DUBm complex |
| Component | GO:0110165 | Cellular anatomical entity |
| Component | GO:0140513 | Nuclear protein-containing complex |
| Component | GO:0140535 | Intracellular protein-containing complex |
| Component | GO:1902493 | Acetyltransferase complex |
| Component | GO:1902494 | Catalytic complex |
| Component | GO:1905368 | Peptidase complex |
| Component | GO:1990234 | Transferase complex |
| Function | GO:0003712 | Transcription coregulator activity |
| Function | GO:0003713 | Transcription coactivator activity |
| Function | GO:0005488 | Binding |
| Function | GO:0008270 | Zinc ion binding |
| Function | GO:0030374 | Nuclear receptor coactivator activity |
| Function | GO:0043167 | Ion binding |
| Function | GO:0043169 | Cation binding |
| Function | GO:0046872 | Metal ion binding |
| Function | GO:0046914 | Transition metal ion binding |
| Function | GO:0140110 | Transcription regulator activity |
Expression profile of Atxn7l3 in omics data
| ID | GSE100499 |
| Title | Exploring acupuncture mechanism in preconditioning of IRI |
| Organism | Rattus norvegicus |
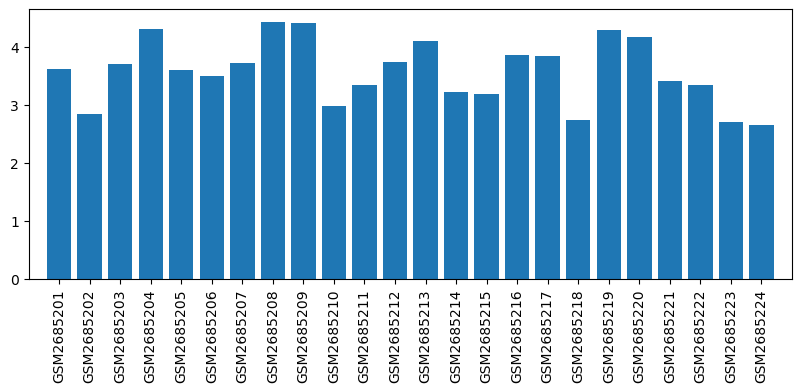
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2685201 | MX1 | tissue: Myocardium gender: male | 3.6157901 |
| GSM2685202 | MX2 | tissue: Myocardium gender: female | 2.8486967 |
| GSM2685203 | MX3 | tissue: Myocardium gender: female | 3.6983094 |
| GSM2685204 | MX4 | tissue: Myocardium gender: male | 4.3118887 |
| GSM2685205 | DZ1 | tissue: Myocardium gender: male | 3.6004355 |
| GSM2685206 | DZ2 | tissue: Myocardium gender: male | 3.5008464 |
| GSM2685207 | DZ3 | tissue: Myocardium gender: female | 3.7216942 |
| GSM2685208 | DZ4 | tissue: Myocardium gender: female | 4.4274564 |
| GSM2685209 | JJ1 | tissue: Myocardium gender: male | 4.4133267 |
| GSM2685210 | JJ2 | tissue: Myocardium gender: female | 2.978376 |
| GSM2685211 | JJ3 | tissue: Myocardium gender: female | 3.335656 |
| GSM2685212 | JJ4 | tissue: Myocardium gender: male | 3.7413216 |
| GSM2685213 | NG1 | tissue: Myocardium gender: male | 4.0926843 |
| GSM2685214 | NG2 | tissue: Myocardium gender: female | 3.2248447 |
| GSM2685215 | NG3 | tissue: Myocardium gender: male | 3.1797411 |
| GSM2685216 | NG4 | tissue: Myocardium gender: female | 3.8522232 |
| GSM2685217 | YL1 | tissue: Myocardium gender: male | 3.8402984 |
| GSM2685218 | YL2 | tissue: Myocardium gender: male | 2.740888 |
| GSM2685219 | YL3 | tissue: Myocardium gender: female | 4.2955084 |
| GSM2685220 | YL4 | tissue: Myocardium gender: female | 4.165705 |
| GSM2685221 | QC1 | tissue: Myocardium gender: male | 3.4153607 |
| GSM2685222 | QC2 | tissue: Myocardium gender: male | 3.3351512 |
| GSM2685223 | QC3 | tissue: Myocardium gender: female | 2.6982627 |
| GSM2685224 | QC4 | tissue: Myocardium gender: female | 2.6572914 |
| ID | GSE136833 |
| Title | Effects of lidocaine on the expression of rat spinal cord |
| Organism | Rattus norvegicus |
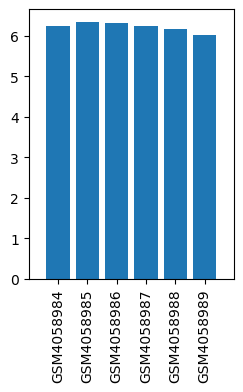
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4058984 | Control_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.240007661 |
| GSM4058985 | Control_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.340838694 |
| GSM4058986 | Control_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.315767698 |
| GSM4058987 | Lidocaine_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.240576905 |
| GSM4058988 | Lidocaine_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.169530304 |
| GSM4058989 | Lidocaine_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.018616404 |
| ID | GSE139220 |
| Title | Expression data from sevoflurane anesthesia treated rat |
| Organism | Rattus norvegicus |
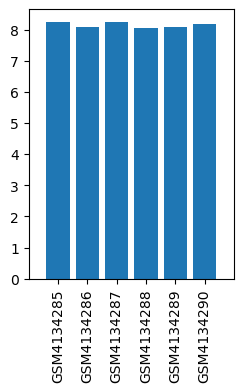
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4134285 | Brain_Control_rep1 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 8.236136 |
| GSM4134286 | Brain_Control_rep2 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 8.077169 |
| GSM4134287 | Brain_Control_rep3 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 8.242316 |
| GSM4134288 | Brain_Sevoflurane_rep1 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 8.045951 |
| GSM4134289 | Brain_Sevoflurane_rep2 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 8.084167 |
| GSM4134290 | Brain_Sevoflurane_rep3 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 8.187292 |
| ID | GSE1616 |
| Title | Genomics of preconditioning |
| Organism | Rattus norvegicus |
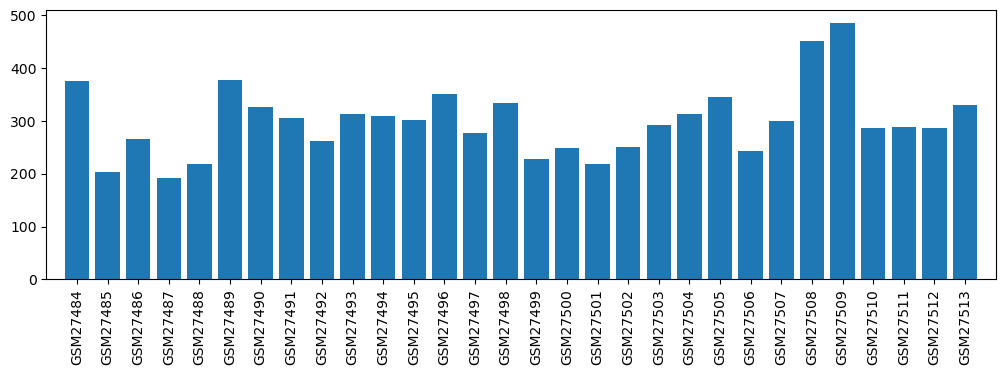
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM27484 | APC_a (anesthetic-preconditioning, 1st replicate) | 375.6 | |
| GSM27485 | APC_b (anesthetic-preconditioning, 2nd replicate) | 203.6 | |
| GSM27486 | APC_c (anesthetic-preconditioning, 3rd replicate) | 266.4 | |
| GSM27487 | APC_d (anesthetic-preconditioning, 4th replicate) | 191.5 | |
| GSM27488 | APC_e (anesthetic-preconditioning, 5th replicate) | 218.4 | |
| GSM27489 | APC_TRI_a (trigger of anesthetic-preconditioning, 1st replicate) | 377.9 | |
| GSM27490 | APC_TRI_b (trigger of anesthetic-preconditioning, 2nd replicate) | 326.1 | |
| GSM27491 | APC_TRI_c (trigger of anesthetic-preconditioning, 3rd replicate) | 305.3 | |
| GSM27492 | APC_TRI_d (trigger of anesthetic-preconditioning, 4th replicate) | 261.4 | |
| GSM27493 | APC_TRI_e (trigger of anesthetic-preconditioning, 5th replicate) | 312.8 | |
| GSM27494 | CTL_a (control, 1st replicate) | 308.9 | |
| GSM27495 | CTL_b (control, 2nd replicate) | 301.3 | |
| GSM27496 | CTL_c (control, 3rd replicate) | 351.8 | |
| GSM27497 | CTL_d (control, 4th replicate) | 277.3 | |
| GSM27498 | CTL_e (control, 5th replicate) | 334.9 | |
| GSM27499 | IPC_a (ischemic preconditioning, 1st replicate) | 228.6 | |
| GSM27500 | IPC_b (ischemic preconditioning, 2nd replicate) | 249.0 | |
| GSM27501 | IPC_c (ischemic preconditioning, 3rd replicate) | 218.0 | |
| GSM27502 | IPC_d (ischemic preconditioning, 4th replicate) | 249.8 | |
| GSM27503 | IPC_e (ischemic preconditioning, 5th replicate) | 292.7 | |
| GSM27504 | IPC_TRI_a (trigger of ischemic preconditioning, 1st replicate) | 313.1 | |
| GSM27505 | IPC_TRI_b (trigger of ischemic preconditioning, 2nd replicate) | 346.0 | |
| GSM27506 | IPC_TRI_c (trigger of ischemic preconditioning, 3rd replicate) | 242.4 | |
| GSM27507 | IPC_TRI_d (trigger of ischemic preconditioning, 4th replicate) | 299.4 | |
| GSM27508 | IPC_TRI_e (trigger of ischemic preconditioning, 5th replicate) | 450.5 | |
| GSM27509 | ISCH_a (ischemia, 1st replicate) | 486.0 | |
| GSM27510 | ISCH_b (ischemia, 2nd replicate) | 286.6 | |
| GSM27511 | ISCH_c (ischemia, 3rd replicate) | 288.7 | |
| GSM27512 | ISCH_d (ischemia, 4th replicate) | 287.5 | |
| GSM27513 | ISCH_e (ischemia, 5th replicate) | 330.9 |
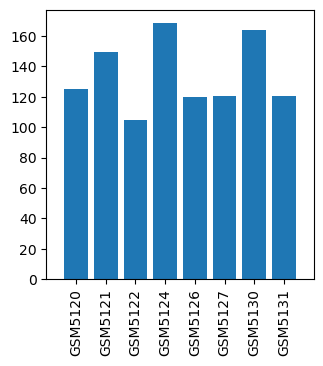
| ID | GSE359 |
| Title | Isoflurane Single Exposures |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5120 | Isoflurane single exposure control-1 | 125.3 | |
| GSM5121 | Isoflurane single exposure control-2 | 149.8 | |
| GSM5122 | Isoflurane single exposure control-3 | 104.7 | |
| GSM5124 | Isoflurane single exposure 1 hr-1 | 168.8 | |
| GSM5126 | Isoflurane single exposure 1 hr-2 | 120.0 | |
| GSM5127 | Isoflurane single exposure 2 hr-1 | 120.7 | |
| GSM5130 | Isoflurane single exposure 2 hr-2 | 163.7 | |
| GSM5131 | Isoflurane single exposure 2 hr-3 | 120.7 |
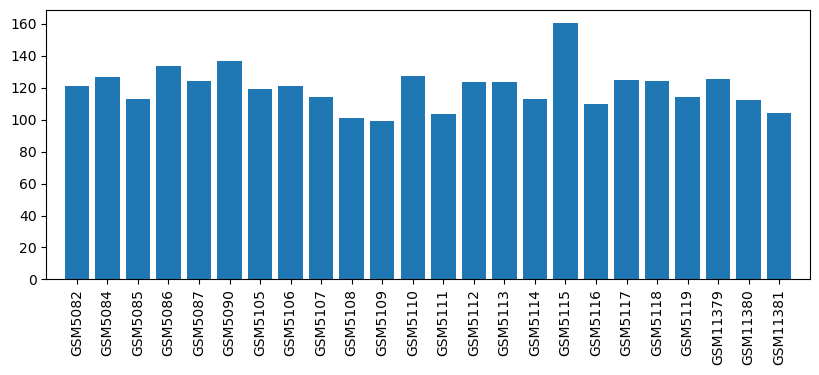
| ID | GSE752 |
| Title | Halothane/Isoflurane Repetitive Exposures |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5082 | Control 1 | 121.4 | |
| GSM5084 | Control 2 | 127.0 | |
| GSM5085 | Control 3 | 112.7 | |
| GSM5086 | Control 4 | 133.7 | |
| GSM5087 | Control 5 | 124.3 | |
| GSM5090 | Control 6 | 137.0 | |
| GSM5105 | Control 7 | 119.2 | |
| GSM5106 | Control 8 | 120.8 | |
| GSM5107 | Control 9 | 114.2 | |
| GSM5108 | Halothane 10 exposure-1 | 100.8 | |
| GSM5109 | Halothane 10 exposures-2 | 99.3 | |
| GSM5110 | Halothane 10 exposures-3 | 127.5 | |
| GSM5111 | Halothane 5 exposures-1 | 103.4 | |
| GSM5112 | Halothane 5 exposures-2 | 123.7 | |
| GSM5113 | Halothane 5 exposures-3 | 123.7 | |
| GSM5114 | Isoflurane 10 exposures-1 | 113.0 | |
| GSM5115 | Isoflurane 10 exposures-2 | 160.7 | |
| GSM5116 | Isoflurane 10 exposures-3 | 110.1 | |
| GSM5117 | Isoflurane 5 exposures-1 | 124.6 | |
| GSM5118 | Isoflurane 5 exposures-2 | 124.0 | |
| GSM5119 | Isoflurane 5 exposures-3 | 114.2 | |
| GSM11379 | Control 10 | 125.2 | |
| GSM11380 | Control 11 | 112.5 | |
| GSM11381 | Control 12 | 104.2 |

